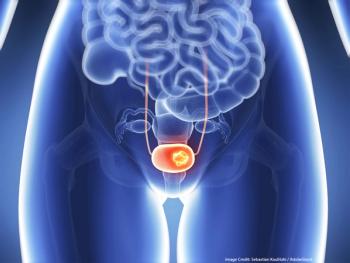
Bladder Cancer Patient on Everolimus Has Complete Response
In an early trial, a patient with bladder cancer has had a 14-month complete response to treatment with the mTOR inhibitor everolimus.
In a phase I trial, a bladder cancer patient has had a 14-month complete response with the combination of the mTOR inhibitor everolimus and pazopanib, a selective multi-targeted receptor tyrosine kinase inhibitor that targets angiogenesis, including the vascular endothelial growth factor (VEGF) receptor. The patient was one of five bladder cancer patients recruited to a clinical trial testing the combination of everolimus and pazopanib in advanced solid tumors.
These results are
Such a so-called ‘exceptional responder,’ one who is acutely sensitive to a specific targeted anticancer or has a particularly durable response, can facilitate a better understanding of the genetic mechanisms underlying the function of these agents, and can improve patient selection as well as clinical trial design. Exceptional responders, as defined by the National Cancer Institute (NCI), are cancer patients who have a partial or complete response to treatment for at least 6 months in a clinical trial in which less than 10% of patients responded.
This type of information can be used to better identify patients who would have a high probability of responding to a particular drug or combination.
“Right now, there is an enormous diversity of how people respond to cancer treatment, from no response to partial to extreme responses. Yet, for most drugs, we don’t know how to predict which patients will respond,” said Nikhil Wagle, MD, of the Dana-Farber Cancer Institute and associate member at the Broad Institute in Cambridge, Mass. This is true even for targeted agents when the patient has a mutation that should make them sensitive to treatment. Detailed analysis of exceptional responders will help identify genomic or other types of markers, and identify salvage drugs and the patients who could take them, said Wagle.
Genetic analysis of the patient’s bladder tumor using whole-exome sequencing revealed two mutations-mTOR E2419K and mTOR E2014K. The constitutively active E2419K mutation has been previously studied in human tissue culture, and also studied in the model organism Saccharomyces cerevisiae, but has not been documented in human tumors. The E2014K mutation is novel and has not been previously described.
Laboratory experiments using cell lines demonstrated that both mutations activated the mTOR pathway, suggesting that they have a relevant biologic role in cancer progression and growth, and create an environment where the tumor is dependent on the active pathway for survival.
The researchers believe that the patient response was a result of everolimus and not pazopanib treatment, as the dose of pazopanib required to overcome the activity of the mTOR mutations is more than 100-fold higher than that for rapamycin, and there were no additional genetic alterations identified that would suggest pazopanib sensitivity.
The current Dana-Farber Cancer Center–initiated trial had enrolled nine advanced, solid tumor patients (five of whom had bladder cancer) who had been previously treated with other anticancer agents. Patients received 1 to 13 cycles of everolimus (5 mg) and pazopanib (400 mg) per day.
Three other patients with bladder cancer on the trial had stable disease for less than 6 months. A patient with adrenal gland cancer had stable disease for 13 months, but none of the lung cancer patients enrolled showed sensitivity to the combination treatment. This combination trial is now being expanded to include a bladder cancer cohort.
While most VEGF receptor tyrosine kinase inhibitors result in significant toxicities in patients, pazopanib has a relatively mild toxicity profile. Pazopanib was combined with everolimus as VEGF receptor inhibition may be complimentary to inhibition of the mTOR pathway: the active mTOR pathway can promote angiogenesis and inhibiting both pathways may lead to better anti-proliferation and anti-angiogenic effects.
Five of nine patients experienced grade 3 or higher including grade 3 febrile neutropenia and thrombocytopenia.
Compared to other tumor types such as colorectal, breast, and prostate, bladder cancer has been less studied and there are no targeted agents currently approved. According to the NCI, about 72,570 people will be diagnosed with bladder cancer this year and 15,210 will die of the disease.
A previous study, from the Memorial Sloan-Kettering Cancer Center in New York, sequenced the entire genome of a tumor from a patient with advanced bladder cancer who responded to everolimus. The sequencing revealed a loss-of-function mutation in TSC1 (tuberous sclerosis complex 1), a regulator of mTOR pathway activation. The patient, a 73-year old woman, had a durable remission following treatment with everolimus. Three other patients in that study also harbored a TSC1 mutation in their tumors and had partial responses to everolimus, while those without the mutation did not respond, suggesting that this mutation conferred sensitivity to the mTOR inhibitor. The study was
Studies such as this one and the current study are allowing Wagle and colleagues to assemble a catalog of mutations that should specific sensitivity to everolimus and other mTOR inhibitors. “We should be screening for these mutations in advance to highlight the population of patients who should receive these drugs,” said Wagle.
These types of studies are already ongoing. There are now basket trials recruiting patients based on specific mutations rather than a tumor type. “There are trials incorporating the identified TSC1 mutations into mTOR inhibitor trials, so this knowledge is already making its way into clinical trials,” said Wagle. “My hope is that we add these new mTOR mutations to the catalog as well.” Information from larger scale genomic efforts that sequence whole-genomes, such as The Cancer Genome Atlas effort, are also providing information on mutations to target.