Might Computers Predict Patient Survival Better Than Pathologists?
Automated computer analysis of pathology slide images can more accurately predict lung cancer patient prognosis than can human pathologists, according to researchers at the Stanford University School of Medicine.
Automated computer analysis of pathology slide images can more accurately predict lung cancer patient prognosis than can human pathologists, according to researchers at the Stanford University School of Medicine in Stanford, Calif. The study was recently
The computers “can assess even tiny differences across thousands of samples many times more accurately and rapidly than a human,” said senior study author Michael Snyder, PhD, in a Stanford
The same approach should work with other cancer types, the authors believe.
“Pathology as it is practiced now is very subjective,” noted Dr. Snyder. “Two highly skilled pathologists assessing the same slide will agree only about 60 percent of the time. This approach replaces this subjectivity with sophisticated, quantitative measurements that we feel are likely to improve patient outcomes.”
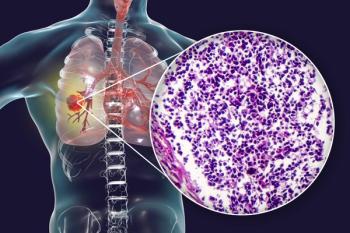
Using machine-learning methods, the research team identified 9,879 disease-related visual features from more than 2,000 hematoxylin and eosin (H&E)-stained histopathology whole-slide images archived in The Cancer Genome Atlas (TCGA). Disease-associated image features included, for example, cell size, shape, texture, and nuclei pixel intensities.
Using small subsets of “top features” these quantitative image features, computer analyses distinguished between stage I adenocarcinoma and squamous cell carcinoma non-small cell lung cancer (NSCLC), and between tumor tissue and adjacent noncancerous tissue exhibiting inflammation, atelectasis or lymphocytic infiltration. Automated analyses also differentiated patients who had short- (<5-year) and long-term (>10-year) survival times after diagnosis-something human observers find difficult using H&E slides alone, the authors noted.
The research team then validated these findings using 294 additional slide images from an independent data set, the Stanford Tissue Microarray (TMA) Database.
Tumor stage and grade offer “limited predictive values” for survival outcomes among patients diagnosed with squamous cell carcinoma, the authors noted.
Because machine learning identified new histopathological visual markers that correlate with survival time, such software might offer new insights into the molecular mechanisms involved in tumor progression, the authors suggested.
However, they also cautioned that slide images submitted to TCGA and the TMA Database might be biased toward more definitive morphological appearances than those seen in “day-to-day practice.”
Newsletter
Stay up to date on recent advances in the multidisciplinary approach to cancer.